 .
.
A cellular automaton (or CA) is a manifold of computing cells which repeatedly update their internal states. The update process is characterized by parallelism, homogeneity, and locality. Parallelism means that all the cells are updated at the same time, in lock-step synchronization. Homogeneity says all cells use the same update rule; and locality says cells only gather information from their nearby neighbors.
Much of the mathematical theory of cellular automata focuses on the large-scale dynamical behaviors of classes of CAs; this involves looking at many different runs of CAs. As a practical matter, it used to be hard to get large CA computations to run rapidly, so mathematicians have mostly looked at CAs in which the cells have only a small amount of information --- for instance, the famous Game of Life CA uses only one bit of internal state per cell. There is, however, no intrinsic reason why CAs should not have state values characterized as one or even several continuous-valued numbers.
Looking at such continuous-valued CAs to some extent duplicates work which numerical analysts have already done under the name of finite difference method simulations. But approaching these numerical simulations from a CA standpoint has the positive effects of (1) incorporating existing CA simulation techniques to achieve a rapidly-running and interactive graphical display, (2) encouraging an artificial life outlook in which emergent simulation properties are examined, and (3) implementing a genetic-algorithm approach whereby a large space of simulation-parameter values can be efficiently explored. The CA framework also provides a conceptual language for discussing a simulation's behavior after instability sets in. Readers with access to a Windows machine may wish to download the CAPOW program [4] in order to assess this for themselves.
For our continuous-valued CAs we let each cell be a small data structure which
contains two floating point numbers that we call U and V. These are to stand for
the intensity and velocity, that is, U stands for u and V stands for 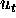 .
.
To carry out this computation we use three buffers, with each buffer being a linear array of cell data structures. The length of these buffers corresponds to the number of space cells being used (or particles being considered). Fermi, Pasta, and Ulam used 64 and 128 cells [2], but we typically use several hundred, up to the horizontal pixel-width of the active screen resolution. At each step of the computation, one buffer can be thought of as holding the current cell values, with the other buffers holding the ``old" cell values and the ``new" cell values. The new buffer is updated on the basis of the values found in the current buffer and the old buffer. Rather than copying values from buffer to buffer, after each update, the names of the buffers are cycled: the new buffer becomes the current buffer, the current buffer becomes the old buffer, and the old buffer becomes the new buffer.
One danger in such continuous-valued simulations is that some of the U or V values may run away to very large or small values which produce a floating point overflow. To avoid this, we add a ``clamping" step to the update process. Some maximum and minimum allowable values of U and V are selected in advance, and whenever a new U or V is computed, it is clamped to lie between the appropriate minimum and maximum; that is, if a value is above the allowed maximum we set it equal to the maximum, and if it is below the allowed minimum we set it equal to the minimum. Clamping is of course a non-physical process, so only a simulation in which no clamping takes place can be regarded as modeling a differential equation.
The values of the U or the V variables in the cells of the current buffer are displayed in two ways: as a graph, and as a space-time diagram. In each case we scale the allowable minimum to maximum range into some small discrete range of possible values. To show a graph, we use the scaled values to measure the vertical displacement of the pixel intended to represent the cell value in question. To represent the cell value as part of a space-time diagram, we use the scaled value to ascertain a color or grey-scale palette index for the pixel representing the cell upon a horizontal line corresponding to that instant of time.
For a given cell at some time j and space position n, 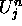 and
and 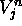 stand
for the respective U and V values of the cell.
stand
for the respective U and V values of the cell. 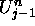 and
and
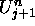 stand for the U values of the cell's left and right neighbors, while
stand for the U values of the cell's left and right neighbors, while
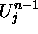 and
and 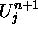 stand for the old and the new values of the cell's U
field.
stand for the old and the new values of the cell's U
field. 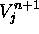 stands for the new value of the cell's V field.
stands for the new value of the cell's V field.
If we track the V values as well as the U values, the scheme corresponding to (23b) is
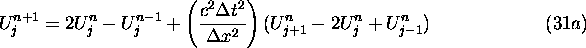

(31a) takes a particularly simple form if c is 1 and

i.e., it becomes
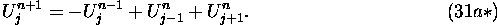
It is typical of the power and economy of cellular automata that such a tiny equation contains the behavior of waves!
One can also think of scheme (31) in an alternate (but equivalent) way which is
perhaps more intuitively obvious: update the velocity by adding
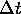 times the acceleration, then update the intensity by adding
times the acceleration, then update the intensity by adding 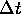 times the
velocity, i.e.,
times the
velocity, i.e.,
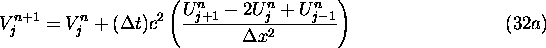

A conceptual advantage of scheme (32) is that it suggests how to expand the scheme to different kinds of accelerating forces. In general if F is a numerical representation of the acceleration caused by a force, then this scheme implies that appropriate schemes will be of the form


In computational practice, the schemes (31) works better than (32) because it is
more ``balanced." That is, we generally are going to be working with small values of
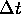 and
and 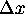 which are similar in size. In (31a), we use them in the form
which are similar in size. In (31a), we use them in the form
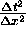 , but in (32a) we use the form
, but in (32a) we use the form 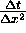 .
Typically the first of these numbers will be near 1, but the second will be quite
large. Multiplying by large numbers reduces a scheme's computational accuracy.
.
Typically the first of these numbers will be near 1, but the second will be quite
large. Multiplying by large numbers reduces a scheme's computational accuracy.
As was mentioned above, unless sufficient care is taken, runaway values of U and V can occur. We discussed how to use a clamping technique to prevent these runaway values from crashing the simulation, but for any physically meaningful simulation, runaway values should not arise in the first place. An analysis of our numerical methods for the above algorithms reveals that runaway values are the result of 1) instability caused by choosing the space and time steps so as to violate the CFL condition or 2) inaccuracy caused by seeding the CA with a clearly discontinuous function.
Regarding the first issue, we recall the CFL stability condition from section 2:

This condition has implications on the time and space scaling of the model.
Since the values of 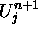 depend only on information in the cell,
depend only on information in the cell, 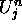 , its
two nearest neighbors,
, its
two nearest neighbors, 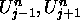 , and its past state,
, and its past state, 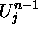 , the
fastest speed at which information can propagate is one space
cell per one time update. So in terms of a cell metric, the maximum
wave speed
c is always 1. In other metrics we can define the maximum speed
c to take
any value as long as we choose an appropriate scale for how we
measure distance
on the
x and t axes. If a horizontal screen inch represents
X spatial units, then a vertical screen inch will represent
, the
fastest speed at which information can propagate is one space
cell per one time update. So in terms of a cell metric, the maximum
wave speed
c is always 1. In other metrics we can define the maximum speed
c to take
any value as long as we choose an appropriate scale for how we
measure distance
on the
x and t axes. If a horizontal screen inch represents
X spatial units, then a vertical screen inch will represent
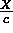 time units. Thus if we want c to be the speed of
light
(roughly
time units. Thus if we want c to be the speed of
light
(roughly 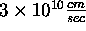 ), and we want the distance
scale to be
natural, with one horizontal screen centimeter representing one
centimeter of x distance, then we must say that one vertical screen
centimeter
represents no more than (roughly)
), and we want the distance
scale to be
natural, with one horizontal screen centimeter representing one
centimeter of x distance, then we must say that one vertical screen
centimeter
represents no more than (roughly) 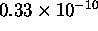 seconds.
seconds.
Regarding the second cause of runaway values, note that if you look
at equation
(31a*), you see that space and time behave like a red and black
checkerboard
of odd and even cells. That is, the values in the ``black" cells
depend only on
the values in the other ``black" cells, while the ``red" cells depend
only on the
other ``red" cells. This means that it is entirely possible for
completely
different patterns to be evolving in the odd and the even space-time
cells. In
the case where 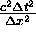 is strictly less than
1.0, there
will be some averaging of information between the black and the red
cells, but
even then the disassociation of the odd-cell and even-cell
simulations can make
itself evident. (We note that in [10], Hayes analyzes this checkerboard effect
for simulations of Burgers's Nonlinear Wave Equation). Attempts to recover a solution
by blending or averaging the values from the two patterns is unwarranted and,
unsurprisingly, leads to bizarre, nonphysical behavior.
is strictly less than
1.0, there
will be some averaging of information between the black and the red
cells, but
even then the disassociation of the odd-cell and even-cell
simulations can make
itself evident. (We note that in [10], Hayes analyzes this checkerboard effect
for simulations of Burgers's Nonlinear Wave Equation). Attempts to recover a solution
by blending or averaging the values from the two patterns is unwarranted and,
unsurprisingly, leads to bizarre, nonphysical behavior.
When you seed this simulation with a discontinuous initial profile, then the
numerical scheme does not simulate the underlying wave equation, since the Taylor
series expansion for u used to derive the computational scheme in section 3 is, of
course, not valid at discontinuities. If the CA is seeded discontinuously, it is
entirely likely that the odd and even cells will behave differently in the resulting
pattern. In order to obtain a range of continuous results we can use initial
shapes for 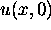 such as a simple sine wave, a Fourier sum of several sine waves,
or we can start with any random smooth pattern. If a simple
discontinuity is introduced into one of our simulations, one sees the expected
``Gibbs ringing" phenomenon whereby high frequency wave components form around the
discontinuity. When the CFL quotient is near unity, the simulation becomes very
sensitive to added discontinuities, as will be discussed in the next section.
such as a simple sine wave, a Fourier sum of several sine waves,
or we can start with any random smooth pattern. If a simple
discontinuity is introduced into one of our simulations, one sees the expected
``Gibbs ringing" phenomenon whereby high frequency wave components form around the
discontinuity. When the CFL quotient is near unity, the simulation becomes very
sensitive to added discontinuities, as will be discussed in the next section.
For the quadratic nonlinear wave equation, we use the scheme in (29),
and set 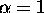 , so that we have
, so that we have
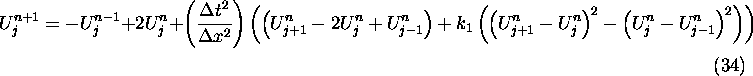
where 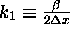 .
In the case of the cubic nonlinear wave equation, (30) gives the
scheme:
.
In the case of the cubic nonlinear wave equation, (30) gives the
scheme:
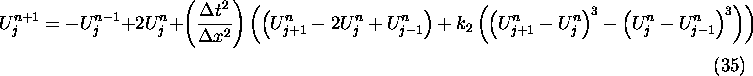
where 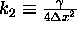 .
.